Welcome to the Baym Lab at Harvard Medical School! We are part of the Departments of Biomedical Informatics and Microbiology, the Harvard Program in Therapeutic Science, and the Broad Institute of Harvard and MIT.
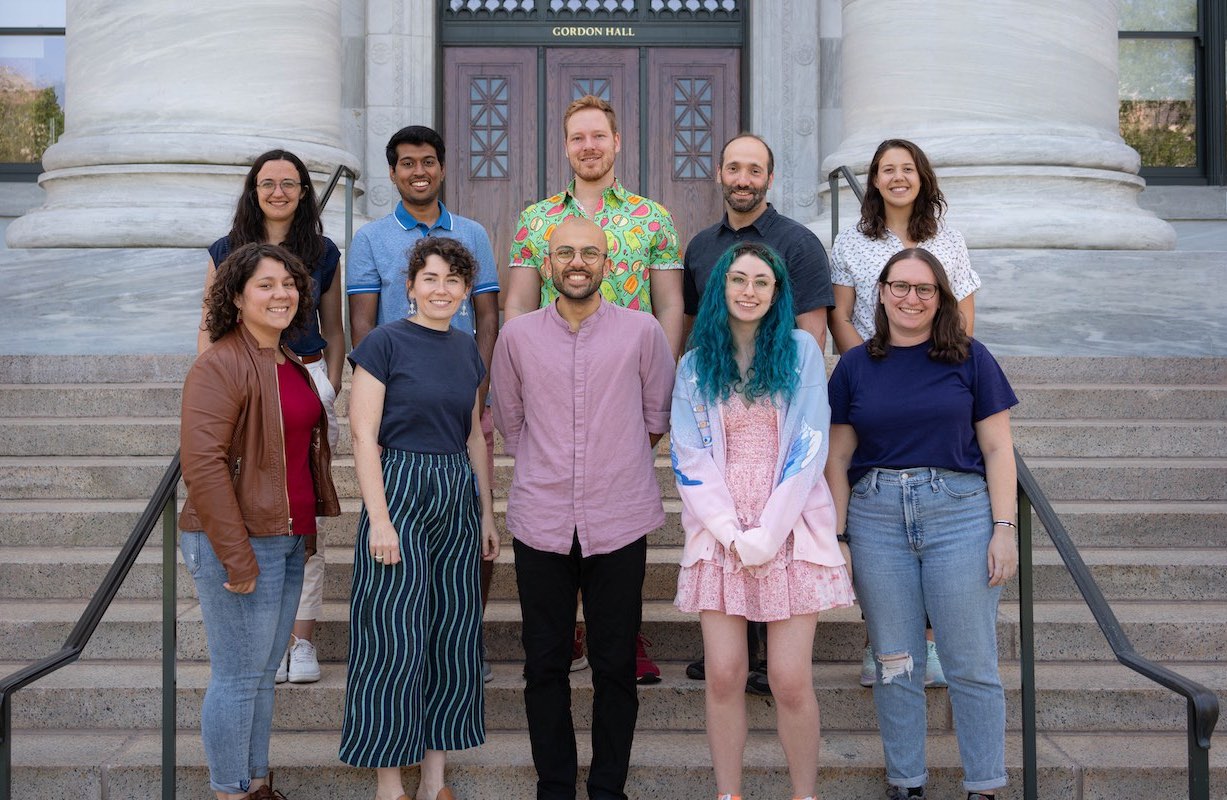 The lab in August 2023
The lab in August 2023
Antibiotic resistance is one of the defining public health threats of our time. Every time a new antibiotic has been introduced, resistance rapidly follows. We study how resistance evolves and more broadly how the mobile genetic elements that allow bacteria genomic flexibility evolve and co-evolve with their hosts. Through this we aim both to better understand microbial evolution, and to discover and develop technologies to study and counter resistance.
Our research ranges from using synthetic systems to study basic evolutionary biology and bacteriology to analysis of clinical sequencing data to technology development.
Or, to describe what we do in Up Goer Five terms:
Our research is supported by the NIH/NIGMS, the NIH/NHGRI, and the NSF.
Lab News
December 2025: Fernando has started a new position a aStaff Scientist in Theoretical Biology at ETH Zürich. Congratulations, Fernando!
November 2025: Fernando's paper, Intracellular competition shapes plasmid population dynamics, is published in ScienceNovember 2025: Kepler and Fernando's paper, RNA-guided nucleases enable a gene drive of insertion sequences in plasmids, is posted as an updated preprint
September 2025: Arya's paper, Genomic resistance in historical clinical isolates increased in frequency and mobility after the age of antibiotics, with Celia and Mische, is published in Microbial Genomics
June 2025: Sam and Hue joined as summer undergrads. Welcome!
May 2025: Ellie's paper, Phage DisCo: targeted discovery of bacteriophages by co-culture, with Natalia, Kesther, Carmen, and Siân, is published in mSystems
May 2025: Our NIH R35, both NSF grants, and Ellie's F31 fellowship were politically terminated.
April 2025: Ellie successfully defended her PhD thesis! Congratulations, Dr. Rand!! 🍾 🎉
April 2025: Karel's paper with Simone and Natalia, Efficient and Robust Search of Microbial Genomes via Phylogenetic Compression, is published in Nature Methods
April 2025: The NIH/NHGRI has awarded a grant for our work on self-evolving molecular barcodes
March 2025: Laura has started as a postdoc. Welcome Laura!
February 2025: Fernando's paper Intracellular competition shapes plasmid population dynamics, is posted as a preprintJanuary 2024: The NIH/NIGMS has awarded our lab a MIRA grant for our work on evolutionary tradeoffs in antibiotic resistance!
January 2025: Michael's position piece with Brad Spellberg and others, Sustainable Solutions to the Continuous Threat of Antimicrobial Resistance, is published in Health Affairs Scholar
January 2025: Arya's paper Genomic resistance in historical clinical isolates increased in frequency and mobility after the age of antibiotics, is posted as a preprint
December 2024: Blox has started as a postdoc. Welcome Blox!
September 2024: Kesther joined the lab as an undergraduate. Welcome, Kesther!
September 2024: Célia has started a new position a an ECDC EUPHEM Fellow at Norwegian Institute of Public Health. Congratulations, Célia!
August 2024: The NSF has awarded a grant for our work on plasmid evolution!
August 2024: Célia's review, From Petri Dishes to Patients to Populations: Scales and Evolutionary Mechanisms Driving Antibiotic Resistance, is published in Annual Review of Microbiology!
August 2024: Siân has started a new position a Research Scientist at the Wadsworth Center and an Assistant Professor at SUNY Albany. Congratulations, Siân!
April 2024: Natalia and Siân's paper, Diverse and abundant phages exploit conjugative plasmids, is published in Nature Communications
March 2024: Shreyas passed his quals. Congratulations, Shreyas!
March 2024: Ellie was awarded an Kirschstein-NRSA/F31 fellowship from the NIH/NIAID for her work on phage discovery!
March 2024: Sophia passed her quals. Congratulations, Sophia!
March 2024: Tatiana joined the lab as a postdoc
March 2024: Michael was promoted to Associate Professor of Biomedical Informatics
February 2024: Natalia successfully defended her PhD thesis! Congratulations, Dr. Quinones-Olvera!! 🍾 🎉
February 2024: Kepler joined the lab as a PhD student
January 2024: Anurag's paper, Changing fitness effects of mutations through long-term bacterial evolution, is published in Science
December 2023: Adele joined the lab as a Master's student
September 2023: Natalia, Fernando, and Celia give talks at the EMBO Plasmids as vehicles of AMR spread meeting
September 2023: Shreyas joined the lab as a PhD student
August 2023: Siân is awarded Best Talk at the Evergreen Phage Meeting!
August 2023: NSF awarded a grant for our work on plasmid-dependent phages!
May 2023: Anurag's paper, Resolving Deleterious and Near-Neutral Effects Requires Different Pooled Fitness Assay Designss, is published in the Journal of Molecular Evolution!
April 2023: Anurag successfully defended his PhD thesis! Congratulations, Dr. Limdi!! 🍾 🎉
April 2023: Sophia joined the lab as a PhD student
March 2023: Siân and Ellie's collaboration with Akos Nyerges and the Church Lab, A swapped genetic code prevents viral infections and gene transfer, is published in Nature
February 2023: Siân's collaboration with Charlie Dulberger and the Rubin and Hatfull Labs, Mycobacterial nucleoid-associated protein Lsr2 is required for productive mycobacteriophage infection, is published in Nature Microbiology
November 2022: Jasmijn's paper Lineage abundance estimation for SARS-CoV-2 in wastewater using transcriptome quantification techniques, is published in Genome Biology
November 2022: Michael receives the Excellence in Mentoring Award from the Systems, Synthetic, and Quantitative Biology PhD program
July 2022: Natalia won the best poster award in the Evolution and Ecology section at Viruses of Microbes 2022!
April 2022: Cristina's paper: Decreased thermal niche breadth as a trade-off of antibiotic resistance, is published in The ISME Journal
March 2022: Amy passed her qualifying exams. Way to be qualified, Amy!
March 2022: Indra has been recognized with a Certificate of Distinction in Teaching for the Fall 2021!
February 2022: Siân's collaboration with the Balskus lab, The bacterial toxin colibactin triggers prophage induction, is published in Nature
January 2022: Célia joined the lab as a postdoc!
January 2022: Karel Brinda has started a new position as INRIA Starting Faculty, at INRIA, Rennes, France. Congratulations, Karel!
December 2021: Indra passed her qualifying exams. Way to be qualified, Indra!
 November 2021: Siân and Natalia's paper appeared on the cover of Cell Host and Microbe!
November 2021: Siân and Natalia's paper appeared on the cover of Cell Host and Microbe!
October 2021: The NASEM consensus report Michael contributed to, Combating Antimicrobial Resistance and Protecting the Miracle of Modern Medicine, has been released
October 2021: Fernando joined the lab as a postdoc!
September 2021: Siân and Natalia's paper: Prophages encode phage-defense systems with cognate self-immunity, is published in Cell Host & Microbe
August 2021: Cristina Herren has started a new position as an Assistant Teaching Professor of Marine and Environmental Sciences at Northeastern University. Congratulations, Cristina!
August 2021: Jasmijn Baaijens has started a new position as an Assistant Professor of Computer Science at TU Delft. Congratulations, Jasmijn!
August 2021: Amy joined the lab as a PhD student
June 2021: Indra joined the lab as a PhD student
April 2021: Karel's paper: Simplitigs as an efficient and scalable representation of de Bruijn graphs, is published in Genome Biology
March 2021: Michael's collaboration with Lee Kennedy-Shaffer and Bill Hanage, Perfect as the enemy of good: tracing transmissions with low-sensitivity tests to mitigate SARS-CoV-2 outbreaks, is published in The Lancet Microbe
March 2021: Michael has received a 2021 A. Clifford Barger Excellence in Mentoring Award from Harvard Medical School
February 2021: Ellie passed her qualifying exams. Way to be qualified, Ellie!
October 2020: Jasmijn is awarded the 2020 BioSB Young Investigator Award! Congratulations Jasmijn!!
October 2020: Arya joined the lab as a PhD student
October 2020: Michael's collaboration with the Zhang Lab at Rice, Metastable hybridization-based DNA information storage to allow rapid and permanent erasure, is published in Nature Communications
August 2020: Ellie joined the lab as a PhD student
July 2020:Siân and Natalia's preprint posted: Prophage-encoded phage defence proteins with cognate self-immunity
June 22, 2020:The wet lab has resumed limited activity with infection control measures.
June 2020: The lab has been awarded a Pew Biomedical Scholarship for 2020-2024! 🎉🎉🎉
June 2020: Victoria and Siân's paper, Forensic microbial system for high-resolution object provenance, is published in Science!
May 9, 2020: Ellie helps the NC Dinos (NC 다이노스) throw out the first pitch.
March 13, 2020:The wet lab closes for Covid-19.
February 2020: The lab has been awarded a Sloan Research Fellowship for 2020-2022! 🎉🎉🎉
February 2020: Karel's paper, Rapid inference of antibiotic resistance and susceptibility by genomic neighbour typing is published in Nature Microbiology!
January 2020: Kaylee joined the lab as a research associate
November 2019: Elle is awarded a Rhodes Scholarship for 2020! Congratulations, Elle!
October 2019: Lucy and Jasmijn join the lab as postdocs
October 2019: Eve's Thesis wins "Best Master Thesis in Life Sciences and Technology" at EPFL! Congratulations, Eve!
September 2019: Siân and Natalia are contributing authors on The fitness landscape of the African Salmonella Typhimurium ST313 strain D23580 reveals unique properties of the pBT1 plasmid in PLOS Pathogens
July 2019: The lab has been awarded a MIRA/R35 Outstanding Early-Stage Investigator Award by NIH/NIGMS for 2019-2024! 🎉🎉🎉
July 2019: Michael contributes to Cell Host and Microbe's Voices on overcoming antibiotic resistance: Sustainable Stewardship Needs Evolution
July 2019: Eve presents a poster at the Microbial Population Biology Gordon Research Conference.
June 2019: Siân presents three talks and two posters at ASM Microbe, and wins a travel award! Wow.
June 2019: Mische joined the lab as a summer undergrad
May 2019: Elle joined the lab as a summer undergrad
March 2019: Natalia passed her quals. Congratulations, Natalia!
February 2019: Anurag passed his quals. Congratulations, Anurag!
February 2019: Cristina speaks at ASLO 2019 in Puerto Rico
October 2018: The lab has been awarded a 2018-2023 Packard Fellowship!
August 28, 2018: Karel's preprint posted: Lineage calling can identify antibiotic resistant clones within minutes
September 2018: Eve joined the lab as a master's student
September 2018: Michael speaks at the Southern California Microbiome Symposium
September 2018: Michael speaks at Lake Arrowhead Microbial Genomics, Siân and Karel present posters
September 2018: Cristina speaks at ISME in Germany
August 2018: Anurag joined the lab as a PhD student
July 2018: Natalia joined the lab as a PhD student
July 2018: Michael speaks at Microbial Stress Response GRC
May 2018: Isabel joined the lab as a summer undergrad
April 2018: Michael speaks at the Harvard MSI Symposium
April 2018: Simone joined the lab as a master's student
April 2018: Siân joined the lab as a postdoc
March 2018: Victoria joined the lab as a technician
February 2018: Scott joined the lab as a visiting postdoc
February 8, 2018: Cristina's preprint posted: Stronger connectivity of the resident gut microbiome lends resistance to invading bacteria
September 2017: Cristina joined the lab
August 2017: Michael speaks at the Microbial Darwinian Medicine Workshop
July 2017: The lab has started! Karel joins the lab as a joint postdoc with the Hanage Lab at HSPH
-->